Article-Genomic Labeling
Spuriously transcribed RNAs from CRISPR-sgRNA expression plasmids scaffold biomolecular condensate formation and hamper accurate genomic imaging
Abstract:
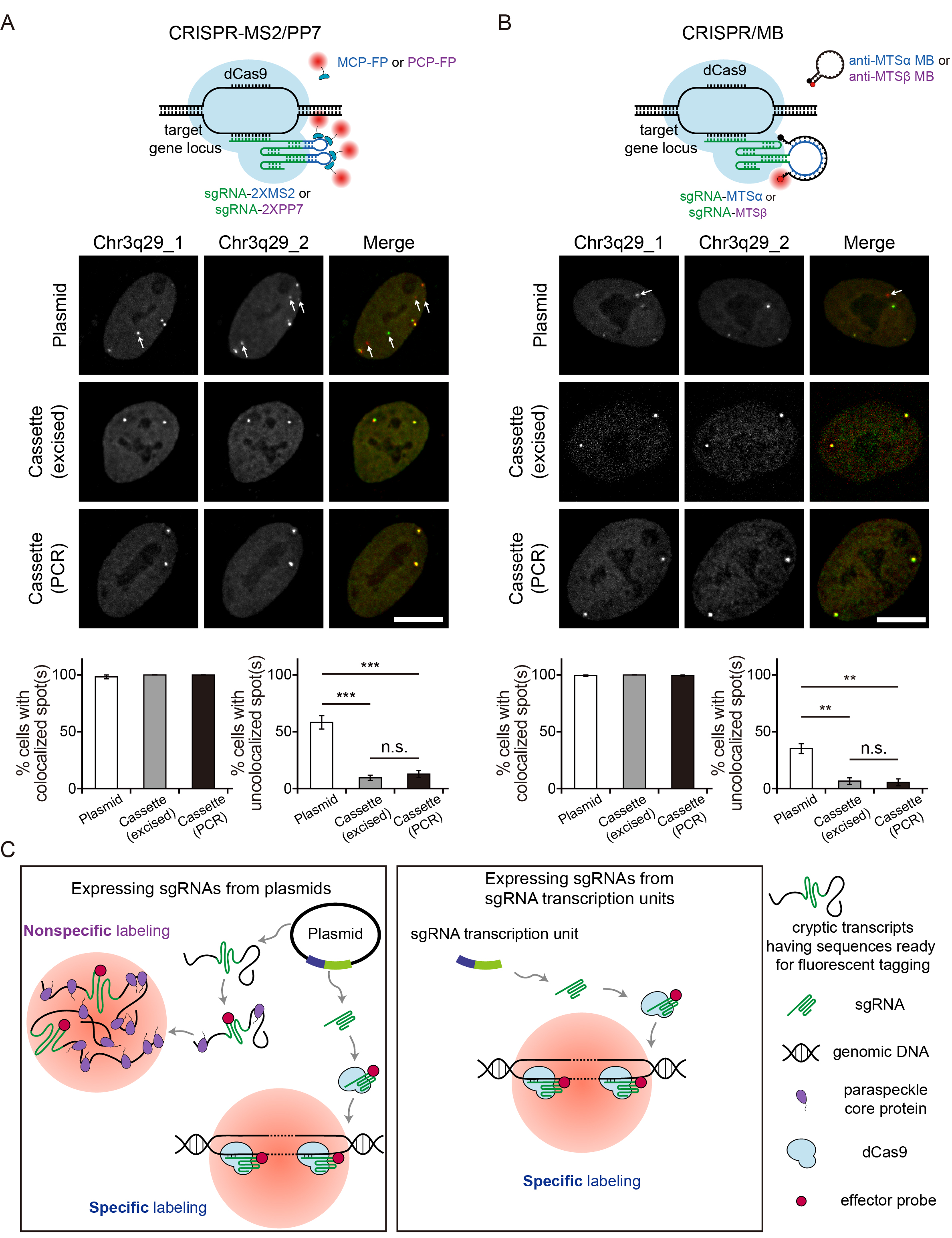
Clustered regularly interspaced short palindromic repeats (CRISPR)-based imaging tools that utilize fluorescently tagged single-guide RNAs (sgRNAs) have enabled versatile analysis of the dynamics of single genomic loci, but the accuracy may be hindered by nonspecific subnuclear probe accumulation, generating false-positive foci in cell nuclei. By examining the subcellular localizations of sgRNA expression plasmids, their RNA transcripts, and several RNA-binding proteins, we found that spuriously transcribed (cryptic) transcripts, produced by sgRNA expression plasmids, are the major contributors of false-positive signals, independent of sgRNA scaffold design or effector probe (i.e. RNA aptamer- or oligonucleotide-based probes) used. These transcripts interact with the paraspeckle core proteins, but not with the sgRNA expression plasmids or the paraspeckle RNA scaffold NEAT1_2, to form nuclear bodies that display liquid-like properties including sphericality, fusion competence, and sensitivity to 1,6-hexanediol. Transfecting sgRNA transcription units (i.e. sgRNA expression cassettes), lacking the plasmid backbones, reduces false-positive signals and enhances genomic imaging accuracy. Overall, this study unveils previously undescribed activities of cryptic plasmid transcripts and presents an easy-to-adapt strategy that can potentially improve the precision of CRISPR-based imaging systems that implement fluorescently tagged sgRNAs.
See full-text: https://doi.org/10.1093/nar/gkaf192
Yachen Ying PUBLICATION
CRISPR imaging Cryptic transcripts Liquid like structure